RNA Biology
1. Analysis of Pathological Nuclear Liquid–Liquid Phase Separation and Drug Discovery
More than 1,000 RNA-binding proteins are known to exist in mammals. These proteins bind to specific RNA sequences and secondary structures (consensus sequences or structures) and regulate the post-transcriptional processing of transcribed RNAs—including alternative splicing, polyadenylation, mRNA stability and metabolism, transport, and translational control—thereby contributing to the diversity of gene expression. This regulatory system is thought to play particularly important roles in highly specialized and structurally complex tissues of higher organisms, such as the nervous system and muscle. However, the overall picture of how these mechanisms function in individual tissues and cell types, and what they comprehensively regulate as a whole, remains far from fully understood. In recent years, increasing attention has been directed toward the possibility that intracellular macromolecules, including RNA and RNA-binding proteins, assemble into liquid-like condensates that function as membrane-less organelles. We focus on abnormalities in this liquid–liquid phase separation and are investigating the potential for drug discovery through compounds that correct such abnormalities. This line of research may ultimately lead to a transformation of conventional drug discovery concepts, which have traditionally been compared to a lock-and-key model.
2. Aberrant RNA Splicing and Disease
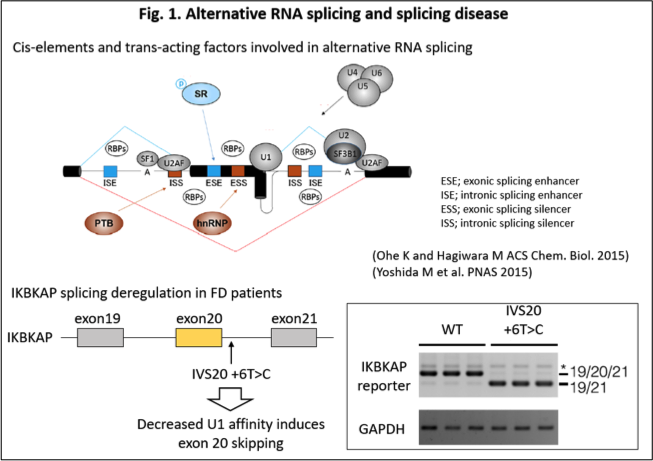
Although RNA-mediated disorders encompass a wide range of conditions, our group has focused particularly on those caused by abnormalities in alternative RNA splicing. In many cases, precursor RNA is regulated through RNA splicing, whereby intronic regions are removed and exonic regions are joined to generate templates for protein translation. This process can occur either constitutively or in a regulated, alternative manner. The former is referred to as constitutive RNA splicing, whereas the latter is known as alternative RNA splicing. Through alternative RNA splicing, multiple isoforms can be generated from a single gene, thereby creating tissue-specific and developmental stage-specific patterns of gene expression and contributing to proteome diversity. By contrast, in certain diseases, including inherited disorders and tumors, abnormalities in RNA splicing caused by mutations in cis-regulatory sequences or alterations in trans-acting factors can become direct causes of disease. One representative example is familial dysautonomia (FD), in which a T-to-C substitution occurs at the sixth nucleotide of the intron downstream of exon 20 in the IKBKAP gene (IVS20+6T>C). This mutation impairs recognition by U1 snRNP, resulting in exon 20 skipping, reduced expression of the gene product IKAP, and consequent abnormalities in tRNA modification (Fig. 1). Likewise, aberrant splicing of tumor suppressor genes can give rise to hereditary cancers. We investigate the dynamic dysregulation of disease-associated splicing events at the individual level and, in parallel, pursue therapeutic strategies based on small-molecule compounds, as described below.
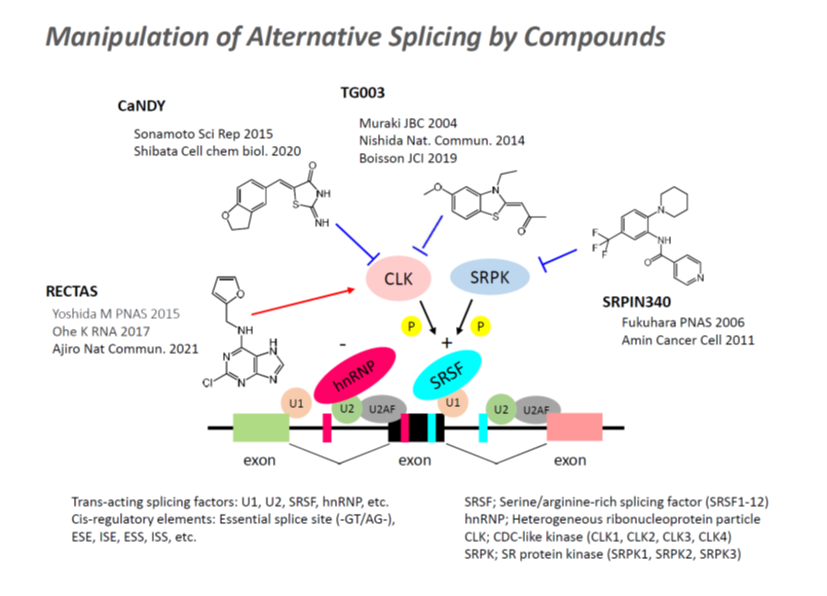
3. Therapeutic Strategies for RNA Splicing Disorders Using Small-Molecule Compounds
(see also the section on transcriptome-based drug discovery)
In seeking to control aberrant splicing in disease, we have focused on kinases that regulate the activity of SR proteins. Among the many molecules involved in splicing regulation as trans-acting factors, SR proteins are well known as promoters of exon inclusion in alternative RNA splicing. SR protein kinase (SRPK), cdc2-like kinase (CLK), and dual-specificity tyrosine(Y)-phosphorylation-regulated kinase (DYRK) promote spliceosome formation for specific splicing events through phosphorylation of SR proteins. Our laboratory has developed the SRPK inhibitor SRPIN340, the CLK inhibitor TG003, DYRK inhibitors, and related analogs (Fukuhara et al., Proc Natl Acad Sci U S A. 2006; Muraki et al., J Biol Chem. 2004; Ogawa et al., Nat Commun. 2010), and has demonstrated the therapeutic potential of regulating SR protein activity in splicing-related diseases. For example, in a subset of Duchenne muscular dystrophy cases, a mutation in exon 30 of the Dystrophin gene (c.4303G>T) generates a premature stop codon. Inhibition of CLK by TG003 induces skipping of exon 30, thereby promoting expression of an isoform that retains dystrophin function and offering a potential therapeutic benefit (Nishida et al., Nat Commun. 2011). In addition, inhibition of SRPK1 by SRPIN340 has been shown to suppress angiogenesis in tumors and the choroid through splicing alterations in the VEGF gene, as well as to inhibit viral RNA function (Dong et al., Mol Vis. 2013; Amin et al., Cancer Cell. 2011; Nowak et al., J Biol Chem. 2010; Fukuhara et al., Proc Natl Acad Sci U S A. 2006).
On the other hand, we have developed RECTAS, a compound that selectively promotes exon inclusion, and found that treatment with RECTAS restores exon 20 inclusion of IKBKAP in familial dysautonomia (Yoshida et al., Proc Natl Acad Sci U S A. 2015; Ajiro et al., Nat Commun. 2021). RECTAS2.0, which was generated through synthetic optimization of RECTAS to enhance efficacy and improve pharmacokinetic properties, corrects aberrant splicing of the GLA gene and reduces Gb3 accumulation in cardiac Fabry disease following oral administration (Awaya et al., Sci Adv. 2025; Kyoto University research news). Because these compounds act through the mechanism illustrated in the figure below, their effects are highly selective and have minimal impact on normal splicing. We believe that harnessing the exon inclusion– and exon skipping–inducing activities of such splicing-modulating compounds will lead to the development of novel therapeutic strategies for RNA splicing–related diseases.